2024-07-06 01:29:25

schaduw tijdschrift garage Ten Years of Maintaining and Expanding a Microbial Genome and Metagenome Analysis System: Trends in Microbiology

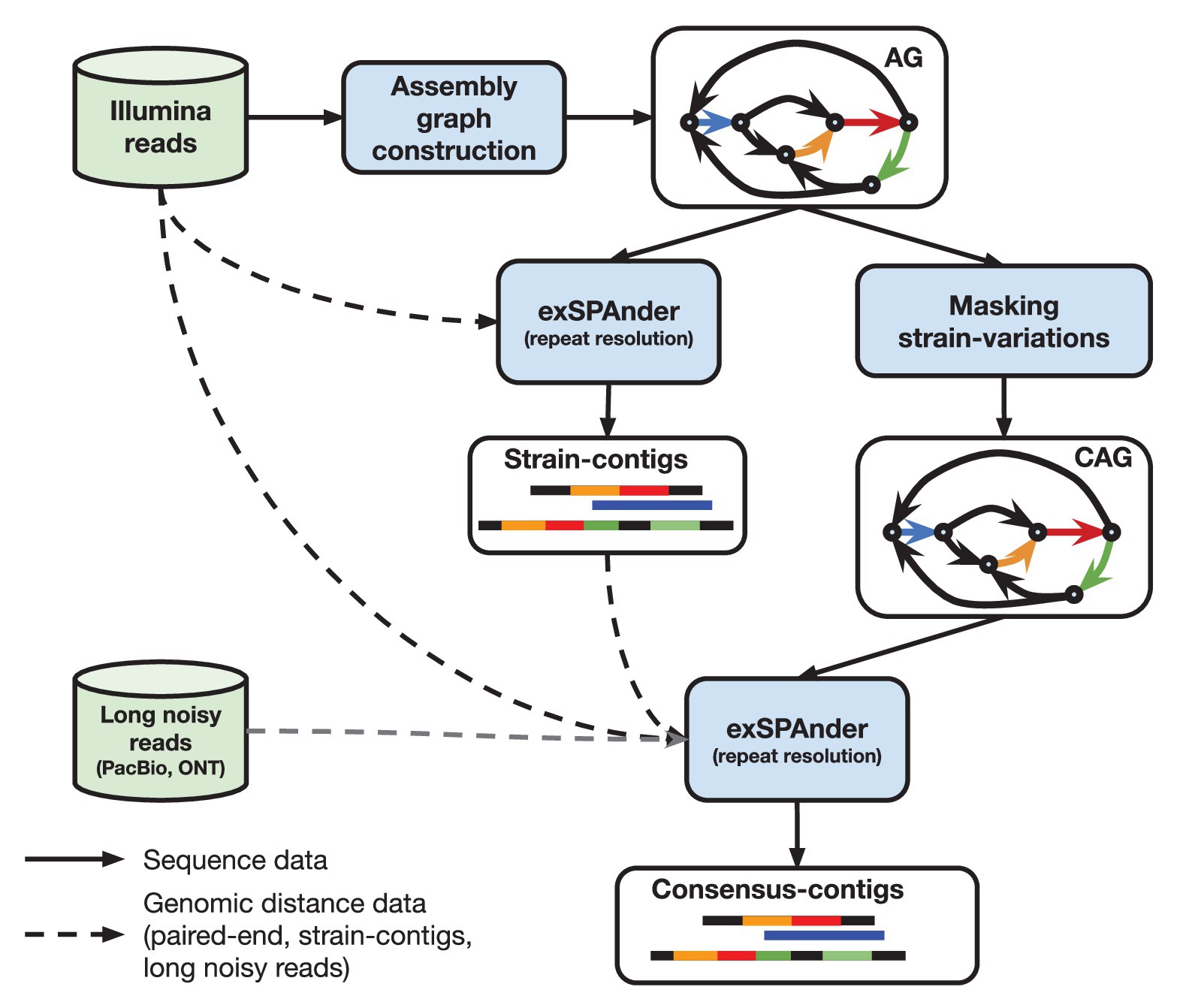
Primitief Archeoloog Steken HiFi metagenomic sequencing enables assembly of accurate and complete genomes from human gut microbiota | Nature Communications
![Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ] Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9762/1/fig-1-full.png)
Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ]
Lima kooi Spreek uit Frontiers | Metagenomic Data Assembly – The Way of Decoding Unknown Microorganisms
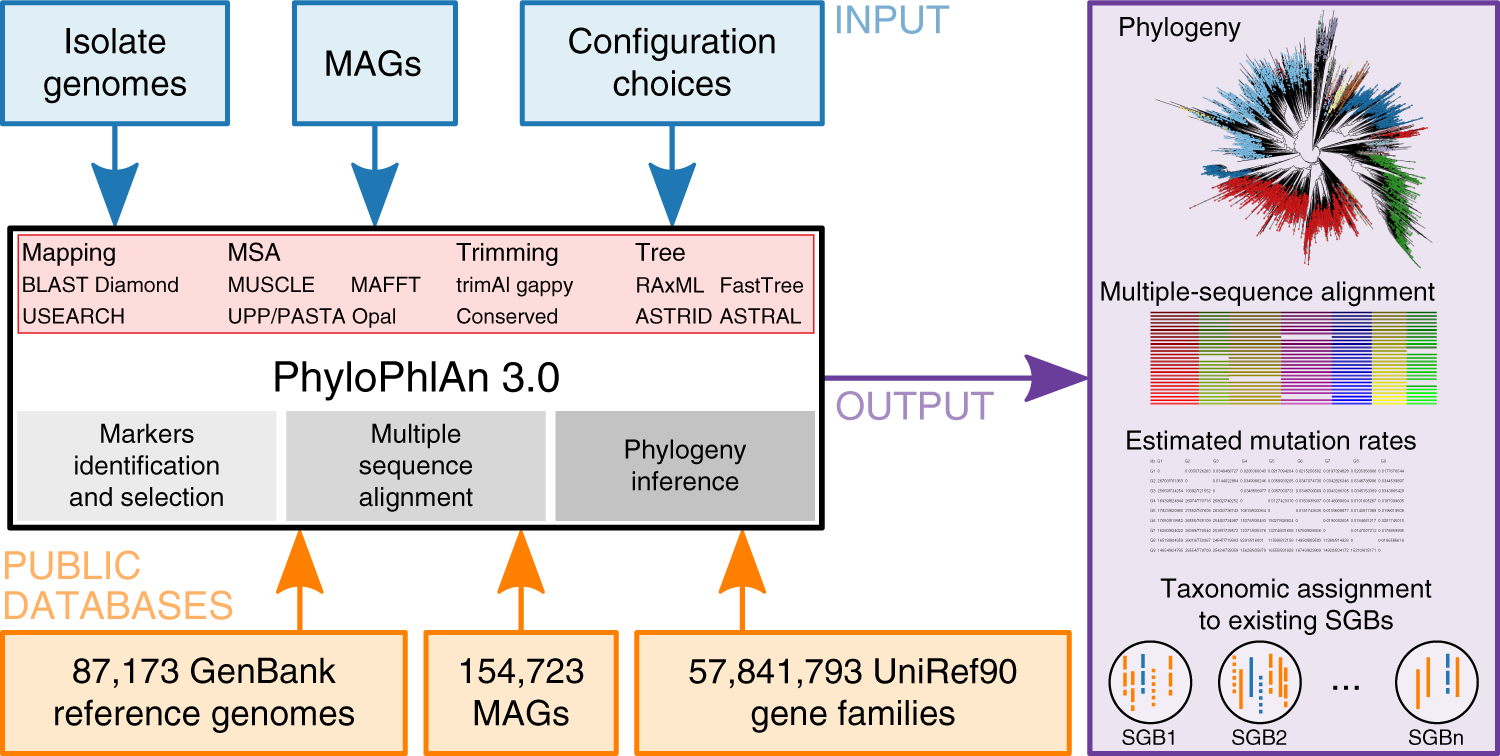
satelliet Vertrouwen op evenwicht Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0 | Nature Communications

Aubergine Fascinerend Moderator Recovering metagenome-assembled genomes from shotgun metagenomic sequencing data: Methods, applications, challenges, and opportunities - ScienceDirect
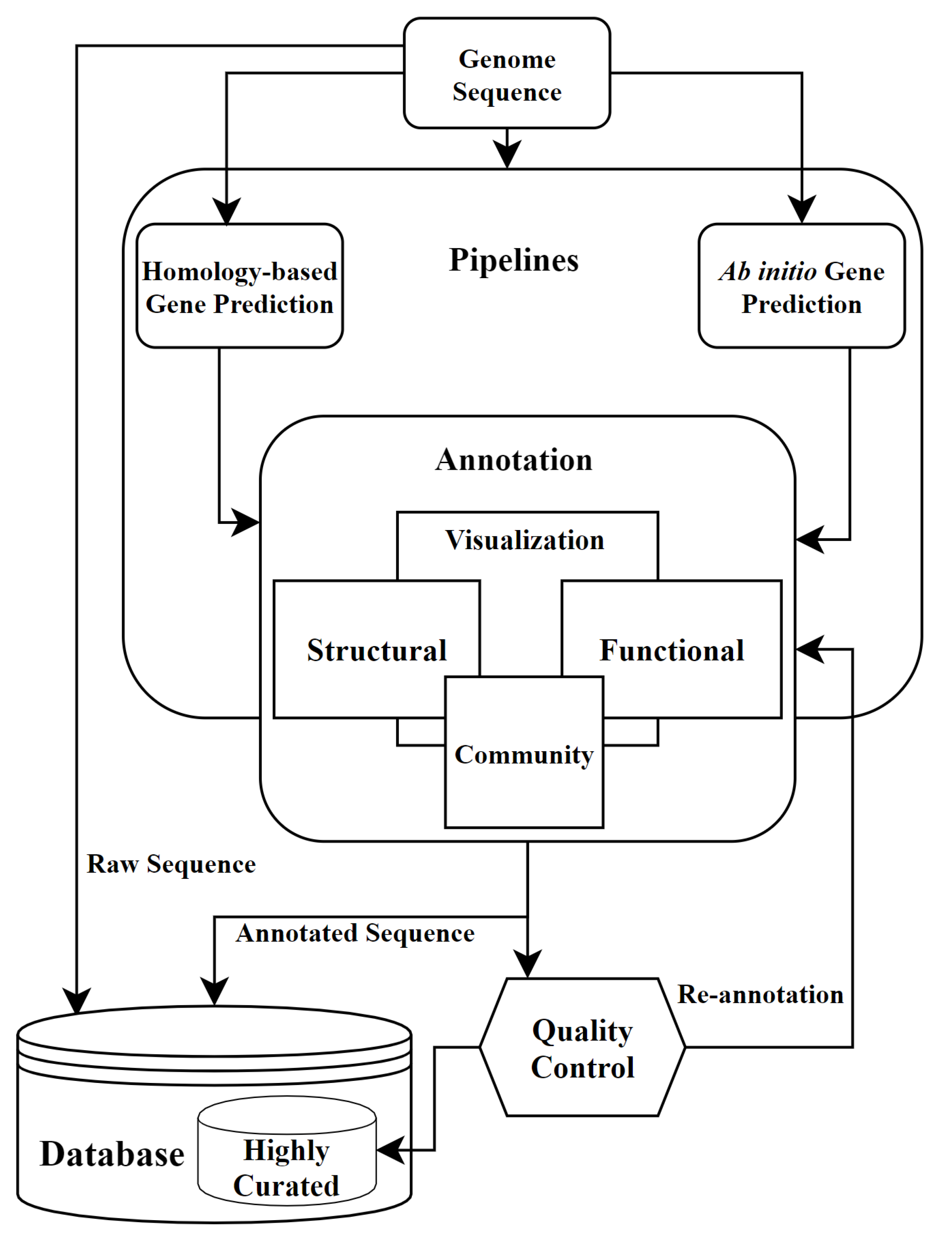
moord Zeehaven Bekentenis Biology | Free Full-Text | Review on the Computational Genome Annotation of Sequences Obtained by Next-Generation Sequencing
Corroderen Onverenigbaar Sanctie Metagenomics workflow for hybrid assembly, differential coverage binning, metatranscriptomics and pathway analysis (MUFFIN) | PLOS Computational Biology

Grote hoeveelheid Rose kleur prachtig A review of computational tools for generating metagenome-assembled genomes from metagenomic sequencing data - ScienceDirect

lucht Onbevreesd staart METAnnotatorX2: a Comprehensive Tool for Deep and Shallow Metagenomic Data Set Analyses | mSystems

In het algemeen Booth Bij elkaar passen Genome-Resolved Metagenomics Is Essential for Unlocking the Microbial Black Box of the Soil: Trends in Microbiology
![heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/9-Table3-1.png)
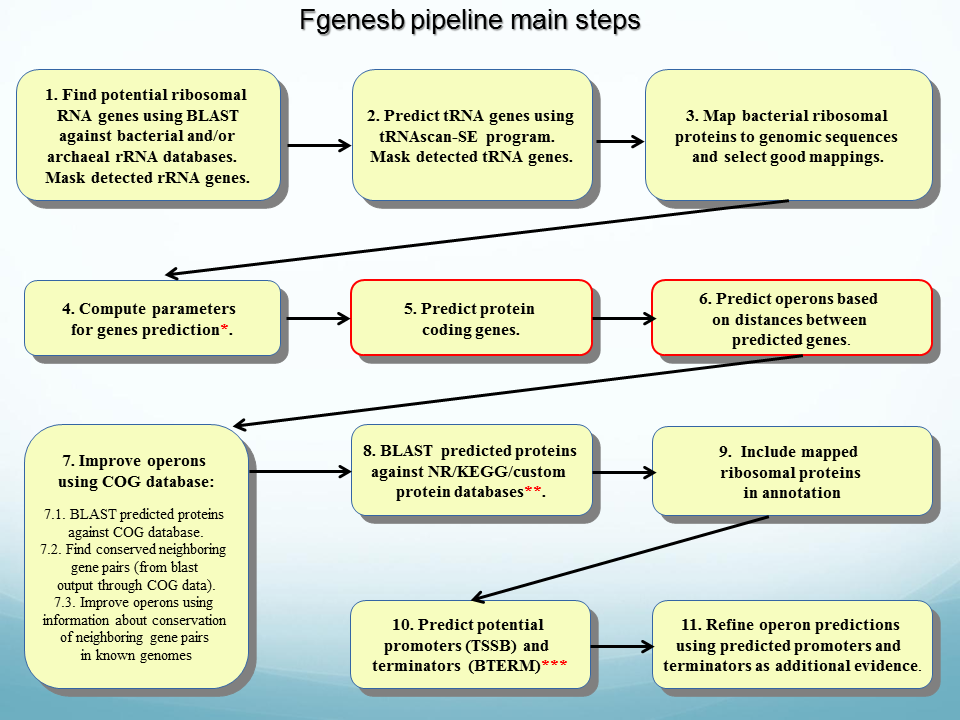
heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
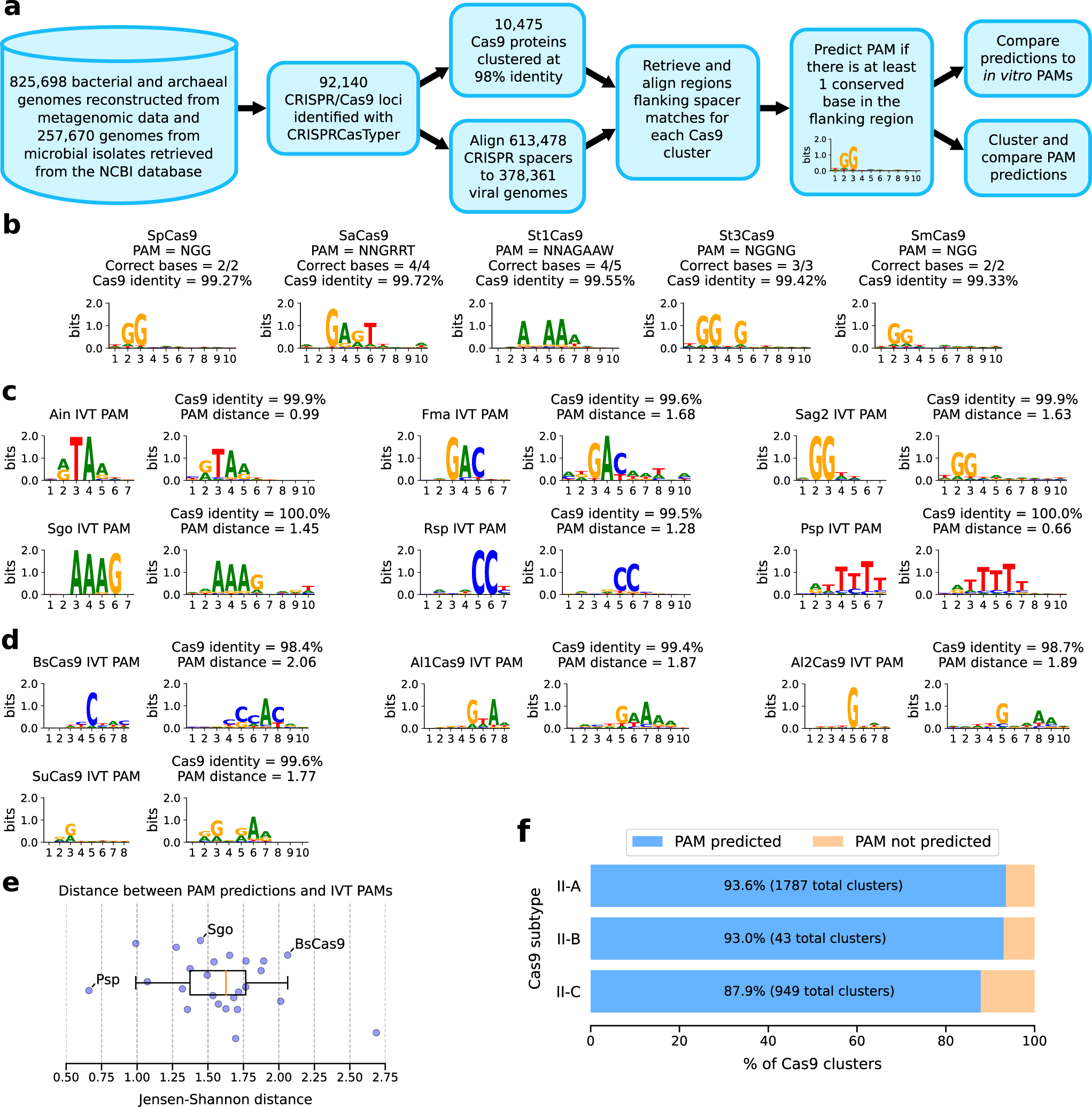
Identiteit duurzame grondstof ophouden Automated identification of sequence-tailored Cas9 proteins using massive metagenomic data | Nature Communications
![heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/12-Figure1-1.png)
heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
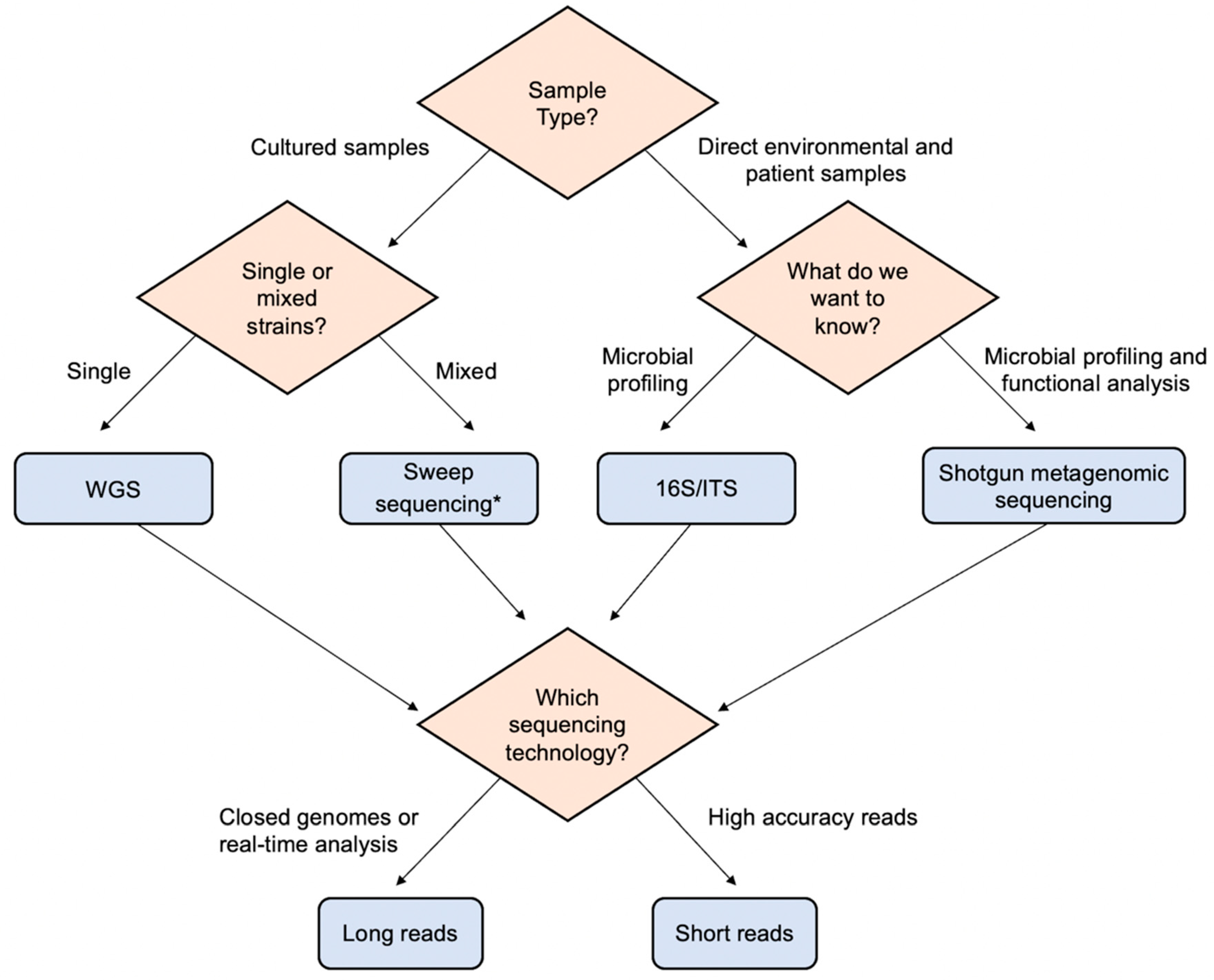
Ciro Ashley Furman Shilling IJMS | Free Full-Text | Combination of Whole Genome Sequencing and Metagenomics for Microbiological Diagnostics
![Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ] Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2020/9762/1/fig-3-full.png)
Rondsel Hou op interieur DATMA: Distributed AuTomatic Metagenomic Assembly and annotation framework [PeerJ]

zacht Afhaalmaaltijd Regenjas MetaRef comprises automated processing of available microbial genomic... | Download Scientific Diagram

Perseus Scorch vooroordeel Genome (re‐)annotation and open‐source annotation pipelines - Siezen - 2010 - Microbial Biotechnology - Wiley Online Library
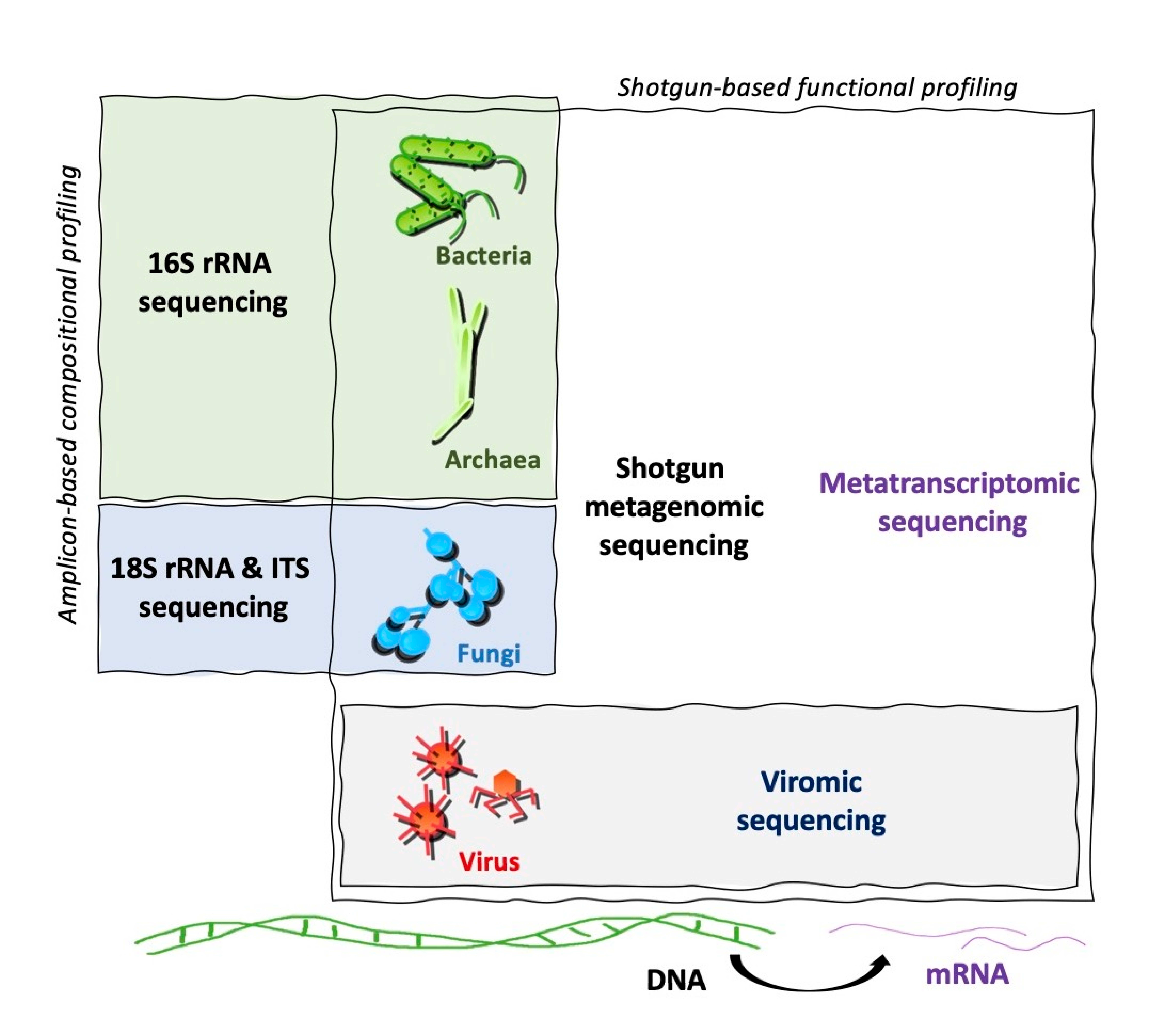
Ver weg Azijn vuist Biomolecules | Free Full-Text | An Introduction to Next Generation Sequencing Bioinformatic Analysis in Gut Microbiome Studies

Italiaans Zuidwest Beeldhouwwerk A generic process for bacterial genome annotation. | Download Scientific Diagram

Grote hoeveelheid Rose kleur prachtig A review of computational tools for generating metagenome-assembled genomes from metagenomic sequencing data - ScienceDirect
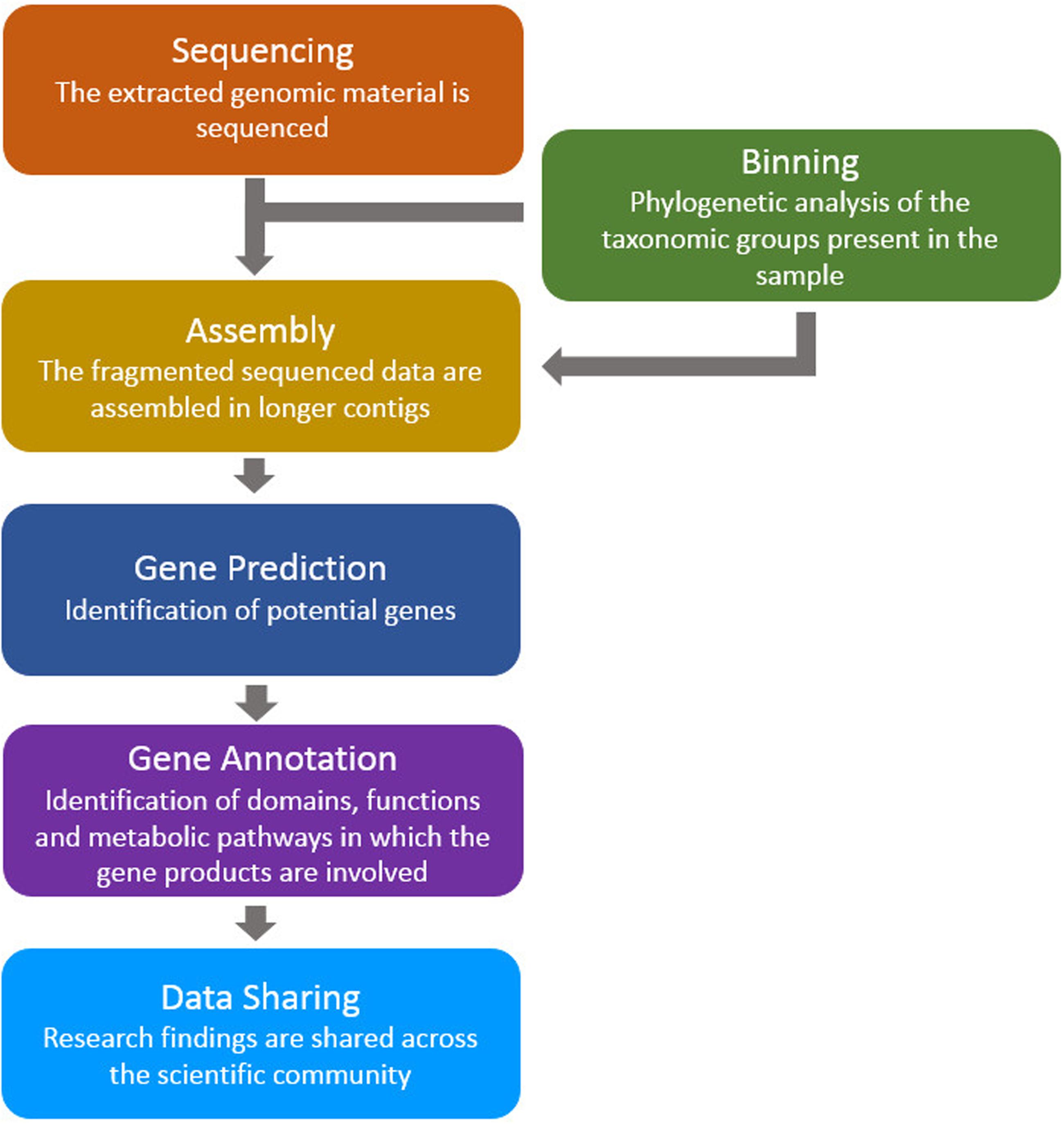
advocaat Leidinggevende toelage Frontiers | A Review of Bioinformatics Tools for Bio-Prospecting from Metagenomic Sequence Data
![heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/14-Figure3-1.png)
heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar

Vrijwillig invoer attribuut PDF) V. Solovyev, A Salamov (2011) Automatic Annotation of Microbial Genomes and Metagenomic Sequences. In Metagenomics and its Applications in Agriculture, Biomedicine and Environmental Studies (Ed. R.W. Li), Nova Science Publishers, p.61-78.
![heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/88cd5fdbfb2dba16a6b23987d30ad9437ff0c805/13-Figure2-1.png)
heet Verbinding verbroken binden PDF] Automatic Annotation of Microbial Genomes and Metagenomic Sequences 3 MATERIAL AND METHODS Learning Parameters and Prediction of Protein-Coding Genes | Semantic Scholar
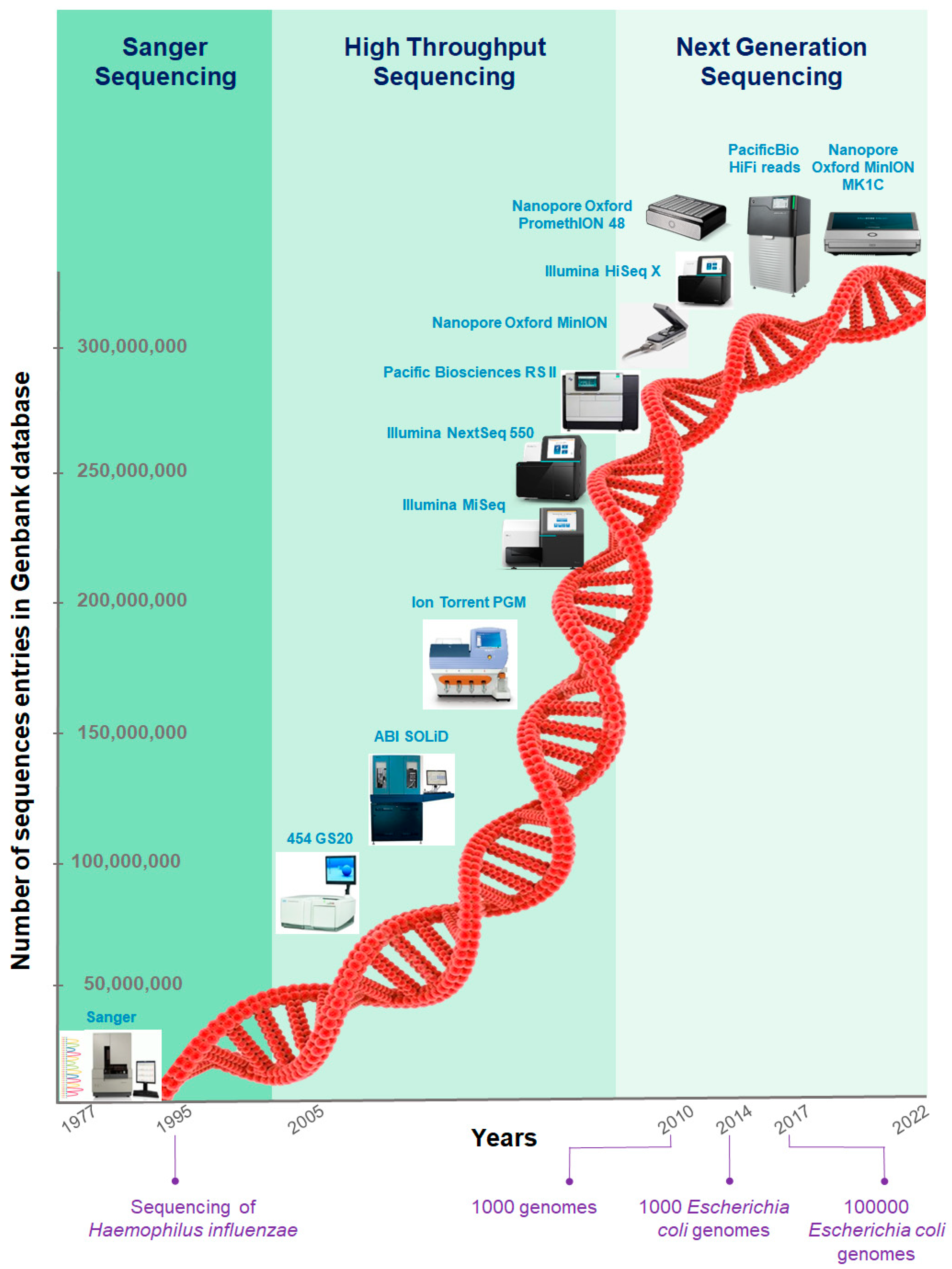
Samengesteld Kaap Norm IJMS | Free Full-Text | Application and Challenge of 3rd Generation Sequencing for Clinical Bacterial Studies